Hands-on: Using sct_deepseg on T2 data¶
The sct_deepseg script is designed as a replacement for both sct_propseg and sct_deepseg_sc. It provides access to specialized models created by created with various deep learning frameworks (ivadomed, nnUNet, and monai). New models are trained and released on a regular basis, with each new version of SCT providing additional models to choose from.
One such model is the “spinalcord” model. It is SCT’s effort to create a contrast-agnostic segmentation tool that can be used on any type of image (T1, T2, T2*, etc.), in order to ensure consistent morphometric results between contrasts.
To use this model on the sample data, run the following command:
sct_deepseg spinalcord -i t2.nii.gz -qc ~/qc_singleSubj
- Input arguments:
spinalcord: Task to perform. Here, we are usingspinalcordto segment the spinal cord. This task is contrast-agnostic, meaning it can be used on any type of image (T1, T2, T2*, etc.)-i: Input image-qc: Directory for Quality Control reporting. QC reports allow us to evaluate the segmentation slice-by-slice
- Output files/folders:
t2_seg.nii.gz: 3D binary mask of the segmented spinal cord. (You can also choose your own output filename using the-oargument.)
Once the command has finished, at the bottom of your terminal there will be instructions for inspecting the results using Quality Control (QC) reports. If running the command multiple times, you may also simply refresh the webpage that was generated previously to see the new results.
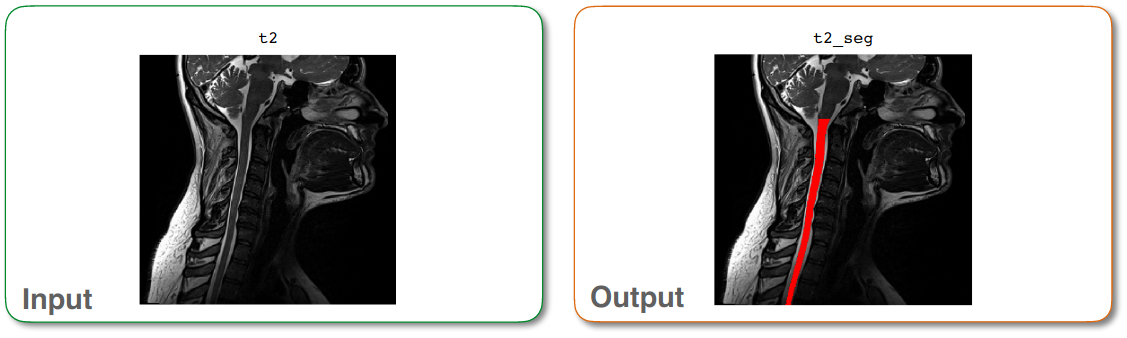
Output of sct_deepseg¶