Lesion segmentation in multiple sclerosis (MS)¶
SCT provides several deep learning-based algorithms to segment lesions in multiple sclerosis (MS). Depending on the image contrast, you can use the following algorithms:
Contrast-agnostic¶
As described in the Contrast-specific vs. contrast-agnostic section, SCT has moved towards developing contrast-agnostic segmentation tools. The lesion_ms model is SCT’s effort to create a contrast-agnostic lesion segmentation tool that can be used on any type of image (T1, T2, T2*, etc.), in order to ensure consistent lesion segmentation results between contrasts.

You can try the lesion_ms on the sample T2w image using the following command:
sct_deepseg lesion_ms -i t2.nii.gz -qc ~/qc_singleSubj
- Input arguments:
lesion_ms: Task-i: Input T2w image with fake lesion-qc: Directory for Quality Control reporting. QC reports allow us to evaluate the segmentation slice-by-slice
MP2RAGE-UNIT1¶
The algorithm lesion_ms_mp2rage was trained on cropped MP2RAGE-UNIT1 images. Details: Cohen-Adad, J., et al. Zenodo release (2023).
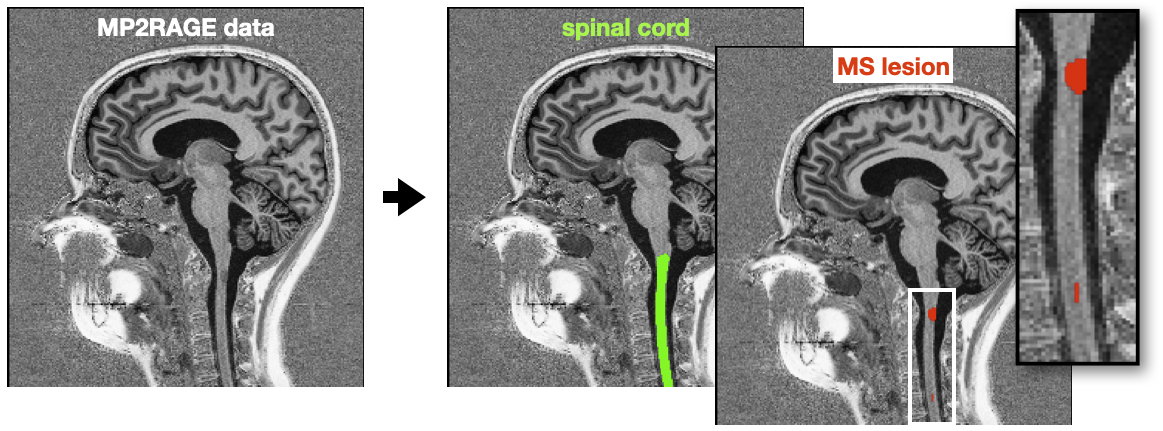
You can try lesion_ms_mp2rage on your own MP2RAGE UNIT1 image using the following commands.
As the model was trained on cropped images, we recommend cropping the input image before running the segmentation.
sct_deepseg spinalcord -i IMAGE_UNIT1 -o IMAGE_seg
sct_crop_image -i IMAGE_UNIT1 -m IMAGE_seg -dilate 30x30x5
sct_deepseg lesion_ms_mp2rage -i IMAGE_UNIT1 -qc ~/qc_singleSubj
- Input arguments:
lesion_ms_mp2rage: Task to perform. In this case, we use thelesion_ms_mp2ragemodel-i: Input MP2RAGE UNIT1 image-qc: Directory for Quality Control reporting. QC reports allow us to evaluate the segmentation slice-by-slice
T2w and T2star¶
The legacy CLI tool sct_deepseg_lesion was trained on T2w and T2star images. Details: Gros, C., et al. NeuroImage (2019).
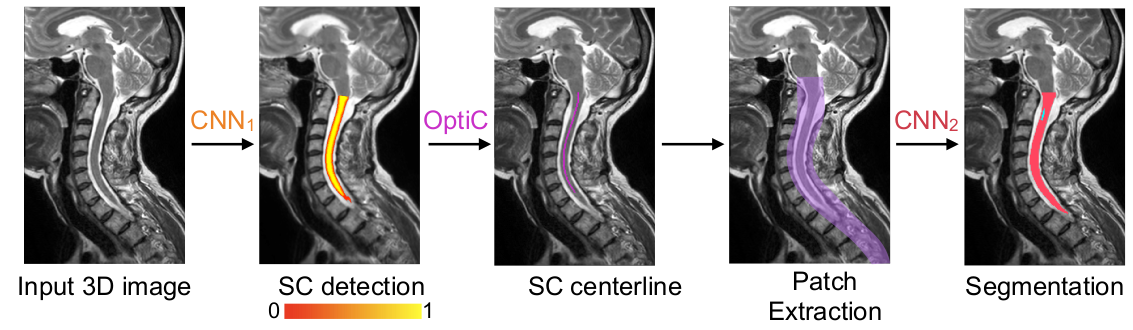
Run the following command to segment the lesion using sct_deepseg_lesion from the input image:
sct_deepseg_lesion -i t2.nii.gz -c t2
- Input arguments:
-i: Input T2w image with fake lesion-c: Contrast of the input image
- Output files/folders:
t2_lesionseg.nii.gz: 3D binary mask of the segmented lesiont2_res_RPI_seg.nii.gz: intermediate segmentation file – you can ignore this filet2_RPI_seg.nii.gz: intermediate segmentation file – you can ignore this file