sct_dmri_separate_b0_and_dwi¶
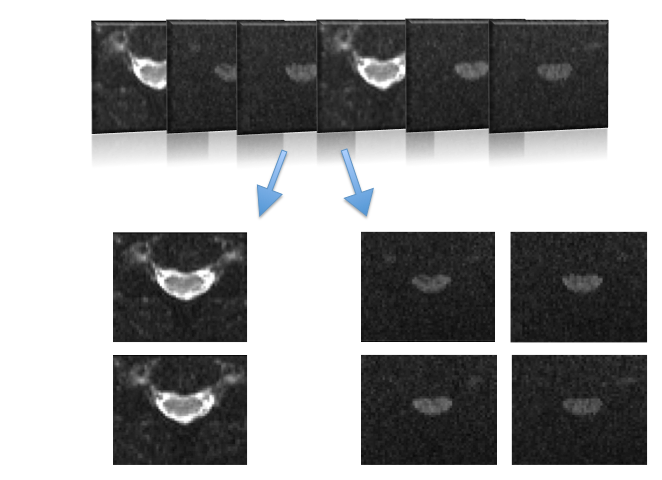
Separate b=0 and DW images from diffusion dataset. The output files will have a suffix (_b0 and _dwi) appended to the input file name.
usage: sct_dmri_separate_b0_and_dwi -i <file> -bvec <file> [-a {0,1}]
[-bval <file>] [-bvalmin <float>]
[-ofolder <folder>] [-h] [-v <int>]
[-r {0,1}]
MANDATORY ARGUMENTS¶
- -i
Diffusion data. Example:
dmri.nii.gz- -bvec
Bvecs file. Example:
bvecs.txt
OPTIONAL ARGUMENTS¶
- -a
Possible choices: 0, 1
Average b=0 and DWI data. 0 = no, 1 = yes
Default:
1- -bval
bvals file. Used to identify low b-values (in case different from 0). Example:
bvals.txtDefault:
''- -bvalmin
B-value threshold (in s/mm2) below which data is considered as b=0. Example:
50.0- -ofolder
Output folder.
Default:
'.'
MISC ARGUMENTS¶
- -v
Possible choices: 0, 1, 2
Verbosity. 0: Display only errors/warnings, 1: Errors/warnings + info messages, 2: Debug mode.
Default:
1- -r
Possible choices: 0, 1
Remove temporary files.
Default:
1